Data and study design
A retrospective study was conducted using EHR data from UF Health IDR, an enterprise data warehouse integrating different patient information systems across the UF Health system. In this study, 125,356 adult patients with 217,221 inpatient encounters who underwent at least one surgical procedure between 2011 and 2022 were initially identified. Patients were excluded if they lacked (1) outcome information (i.e., LOS or mortality), (2) provider information, or (3) recorded surgery dates. After applying the exclusion criteria, the final cohort comprised 136,647 inpatient encounters of 102,768 patients. For this study, the index date was defined as the date of the last surgical procedure within each hospital encounter. Predictors were extracted for each patient during their inpatient stay to the point of discharge, while outcomes were assessed after the index date. The data extraction process is visualized in Fig. 5. This study was conducted in accordance with the ethical principles outlined in the Declaration of Helsinki. This study was approved as exempt by the University of Florida Institutional Review Board (IRB) under protocol number IRB202202634. The research involved the use of de-identified EHR data to ensure participant privacy and confidentiality.
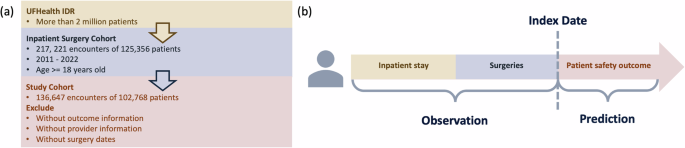
a Study cohort construction process. b Patient timeline. The study cohort consists of adult inpatient surgery patients with documented provider and outcome information. Risk factors are observed during the inpatient stay, prior to the completion of the last surgery. The model is designed to predict patient safety outcomes based on the pre-surgery observation period.
Study outcomes
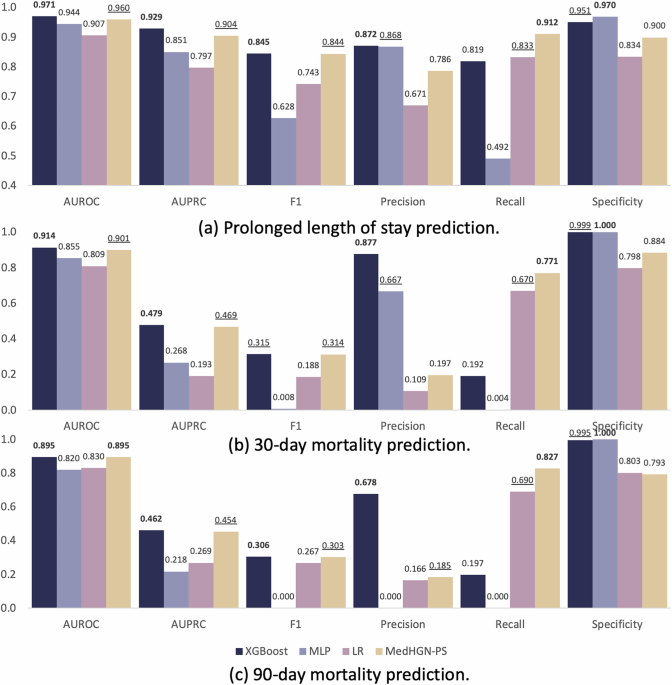
Three study outcomes were validated in this study: PLOS, 30-day and 90-day all-cause mortality after the index date. To determine PLOS, the 75th percentile was used as the cut-off threshold according to the related works37,38,39,40,41. The 30-day and 90-day mortalities are two important criteria set by the Centers for Medicare & Medicaid Services (CMS) for hospitals42.
Covariates
Various patient demographic characteristics recorded at the time of admission were incorporated as predictive variables, including age, weight, height, BMI, race, ethnicity, sex, and marital status.
Clinical factors and calendar-related factors during the patient’s perioperative period were extracted. For clinical factors, comorbidities, the CCI35, the ASA-PS43, surgery services received, procedure interval, anesthesia-related information, and the presence of delirium44 were included. The procedure intervals, including the durations between various operative stages: from room start to induction, from induction to incision, from incision to dressing, from dressing to emergency, from emergency to room end, and duration from the anesthesia start to end were also extracted. Anesthesia-related information was incorporated, including type of anesthesia, and occurrence of anesthesia block. Calendar-related factors, which can potentially impact health quality, were also considered; for example, the type of day (weekday or weekend) may interact with provider shifts, influencing healthcare quality45. These features included the day of week, type of day, admission source, and care unit type.
Preoperative ICU stay has been recognized as a risk factor for adverse outcomes46. Accordingly, records of emergency admissions, preoperative LOS across various care unit types (i.e., intermediate, intensive, and acute), and frequency of transfers between these units were incorporated as predictive features.
A key focus of this study is to discover the impact of care team coordination on outcomes. Therefore, provider information (e.g., expertise type) were included and the interactions among providers within the same encounter and across different encounters were analyzed.
The proposed framework
The proposed analysis framework, MedHG-PS, has three steps: (1) graph construction, (2) GNN modeling, and (3) feature contribution exploration. The inputs to this framework include demographics, perioperative information, transfer records, and provider team details. Before loading inputs to the framework, the raw data was preprocessed by filtering out outliers, standardizing continuous features, and applying one-hot encoding to categorical variables. Missing data were addressed by assigning “Unknown” to missing categorical variables and imputing missing continuous variables using their mean values. To address the imbalanced nature of patient safety outcomes, resampling strategies were further assessed. For each task, data balancing was achieved by either randomly removing negative samples or oversampling positive samples.
A heterogeneous graph \(n=,n\), as shown in Fig. 6a, was first constructed as the starting point using the input variables. The graph consists of three types of nodes \({\mathcal{\in }}{\mathcal{\{}}{\mathcal{E}}{\mathcal{N}}{\mathcal{C}},{\mathcal{P}},{\mathcal{C}}{\mathscr{\}}}\), representing encounter (\({\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\)), provider (\({\mathcal{P}}\)), and care unit (\({\mathcal{C}}\)). All nodes are connected by undirected edges (\({\mathcal{E}}\)), which represent their interaction. For example, if a provider is associated with an encounter, a link is established between the corresponding nodes. Demographic and perioperative information were embedded into a feature matrix \({H}^{{\mathcal{E}}{\mathcal{N}}{\mathcal{C}}}\in {{\mathcal{R}}}^{{\mathcal{E}}{\mathcal{N}}{\mathcal{C}}{\mathcal{\times }}{\mathcal{F}}}\), where \({\mathcal{F}}\) is the number of features. The provider nodes (\({\mathcal{P}}\)) were divided into five subgroups: Surgical Team, Other Clinicians, Nurses, Technicians, and Others. The Surgical Team subgroup includes surgeons, surgical assistants, and surgical fellows/residents. Other Clinicians encompasses providers who do not directly perform surgery, such as anesthesiologists, certified registered nurse anesthetists, anesthesiologist assistants, residents, and physician assistants. The Nurse subgroup includes nurse monitors, nurse practitioners, post-anesthesia care unit nurses, pre-procedural nurses, registered nurses in interventional radiology, and scrub nurses. Technicians consist of cell saver technicians, control room technicians, cardiovascular technicians, gastrointestinal technicians, radiology technologists, surgical technologists, catheterization lab scrubs, and perfusionists. The care unit (\({\mathcal{C}}\)) was classified into three subgroups: Intermediate, Intensive, and Acute. \({\mathcal{P}}\) and \({\mathcal{C}}\) has no feature. For ablation studies, subgraphs only containing \({\mathcal{P}}\) and \({\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\) without \({H}^{{\mathcal{E}}{\mathcal{N}}{\mathcal{C}}}\), \({\mathcal{P}}\) and \({\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\) with \({H}^{{\mathcal{E}}{\mathcal{N}}{\mathcal{C}}}\), \({\mathscr{C}}\) and \({\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\) with \({H}^{{\mathcal{E}}{\mathcal{N}}{\mathcal{C}}}\), was constructed.
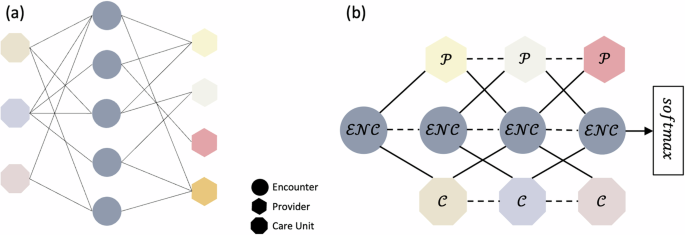
a The representation of the graph. Circles represent encounters, triangles represent providers, and squares represent care units. Different colors represent different providers or care unit types. b MedHG-PS with three stacked layers, where solid edges denote neighbor connections, and dashed edges represent self-relations.
An interpretable and efficient heterogeneous graph convolutional network (ie-HGCN)16 was implemented to model the heterogeneous graph \({\mathcal{G}}\). In the ie-HGCN, given the graph \({\mathcal{G}}\), the task is to learn representation matrices of nodes in \({\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\) for health outcome prediction. The model architecture is illustrated in Fig. 6b, which contains three graph convolutional network (GCN) layers to learn embeddings layer-wise for each node \(\Omega \in {\mathcal{E}}{\mathcal{N}}{\mathcal{C}}\), and then pass to softmax function to generate the outputs. In ie-HGCN, \({H}_{0}^{\Omega }\) refers to the input features, and each GCN layer calculated the node embeddings \({H}_{l}^{\Omega }\) based on previous-layer representation \({H}_{l-1}^{\Omega }\), where \(l\) is the index of layer and \(l\in \left[1,3\right]\), by aggregating self-features \({Z}^{\Omega }\) and neighboring features \({Z}^{\Gamma }\), where \(\Gamma \in {{\mathcal{N}}}^{\Omega }=\{{n|n}\;{\mathscr{\in }}\,{\mathcal{V}}\thinspace{and}\thinspace{n}\;\ne \;\Omega \}\). Since the neighbors of a node in a heterogeneous graph are of different types, the convolution operation cannot be applied directly across all node types simultaneously. Instead, the features \(Z\) was calculated separately by node types. The GCN layer can be formulated as Eq. (1):
$${H}_{l+1}^{\Omega }\,={ELU}\left({a}_{l}^{\Omega }\cdot {Z}_{l}^{\Omega }+\sum _{\Gamma \in {{\mathscr{N}}}^{\Omega }}{a}_{l}^{\Gamma }\cdot {Z}_{l}^{\Gamma }\right)$$
(1)
where \({ELU}(\cdot )\) is the Exponential Linear Unit activation function47. The normalized attention coefficients \({a}^{\Omega }\) and \({a}^{\Gamma }\) for each layer are calculated by Eq. (2). It follows the key, query, value attention mechanism maps a set of queries and a set of key-value pairs to an output. The attention function is implemented as follows:
$$\begin{array}{c}{a}^{\Omega }={{{softmax}}}({ELU}\,([({Z}^{\Omega }{W}_{k}^{\Omega }){{||}}({Z}^{\Omega }{W}_{q}^{\Omega })]\,\cdot {W}_{a}^{\Omega }))\\ {a}^{\Gamma }={{{softmax}}}({ELU}\,([({Z}^{\Gamma }{W}_{k}^{\Gamma }){{||}}({Z}^{\Omega }{W}_{q}^{\Omega })]\,\cdot {W}_{a}^{\Omega }))\end{array}$$
(2)
where trainable matrices \({W}_{k}\) maps \(Z\) into attention keys, and \({W}_{q}\) maps \(Z\) into attention queries. Operation || denotes the row-wise concatenation, and \({W}_{a}^{\Omega }\) is a learnable attention parameter vector. \({Z}^{\Omega }\) and \({Z}^{\Gamma }\) are generated by heterogeneous graph convolution operation. For \({Z}^{\Gamma }\), as is shown in Eq. (3), \({W}^{\Gamma -\Omega }\) along with the row‐normalized adjacency matrix \({\hat{A}}^{\Omega -\Gamma }={{{{diag}}}({A}^{\Omega -\Gamma })}^{-1}\cdot {A}^{\Omega -\Gamma }\) and the previous-layer representation \({H}_{l-1}^{\Gamma }\), together govern the transformation. For \({{Z}}^{\Omega }\), since the \({\hat{A}}^{\Omega -\Gamma }\) are identity matrix, the equation can be simplified to Eq. (4):
$${Z}^{\Gamma }={\hat{A}}^{\Omega -\Gamma }\cdot {H}_{l-1}^{\Gamma }\cdot {W}^{\Gamma -\Omega }$$
(3)
$${Z}^{\Omega }={H}_{l-1}^{\Omega }\cdot {W}^{\Omega -\Omega }$$
(4)
In the final step, the hidden embeddings from the third GNN layer are fed into a softmax function to generate the final prediction probabilities \(\hat{y}\), as shown \(\hat{y}={{\rm{softmax}}}\left({H}_{3}^{\Omega }\,\right)\). During the training phase, the objective is to minimize prediction errors. A gradient-based optimization method is employed to update the trainable parameters, which is illustrated below:
$${{{Loss}}}=\,-\frac{1}{N}\mathop{\sum }\limits_{i=1}^{N}\left[{y}_{{\rm{i}}}log \left({\hat{y}}_{{\rm{i}}}\right)+\left(1\,-\,{y}_{{\rm{i}}}\right)log \left(1\,-\,{\hat{y}}_{{\rm{i}}}\right)\right]$$
(5)
The significance of different input variables influencing patients’ safety outcomes were explored by MPA, SHAP48 analysis and LIME49. MPA assesses the impact of various paths in the graph structure across different node types on patient safety outcomes, offering insights into the most influential clinical interactions. To perform MPA, meta-paths from the graph was constructed by linking different node types. For example, a meta-path could be Encounter -> Provider -> Care unit. For each path \(\Gamma -\Omega\), the mean normalized attention coefficients of edges \({a}_{l}^{\Gamma }\) from each model layer was recorded, capturing the interactions between specific node type pairs (e.g., Acute Care Unit and Encounter nodes). The overall importance of each meta-path was determined by multiplying the attention coefficients along the sequence of edges in the meta-path.
To identify the key demographic and perioperative information influencing patients’ safety outcomes, SHAP48 and LIME, widely used XAI techniques, were employed. SHAP and LIME allow for the interpretation of model predictions by quantifying the contribution of individual features. Given the high dimensionality of the feature space, the Clinical Classifications Software Refined (CCSR) database was applied to aggregate diagnosis and procedure codes into higher-level clinical categories, simplifying the features while preserving their clinical relevance50.
Experimental settings
The proposed analysis was conducted on the University of Florida’s HiperGator high-performance computing cluster ( The experiment used 16 cores of AMD EPYC 7742 processors, 64GB memory, and an Nvidia A100 GPU. For framework implementation and data processing, Python version 3.11 with the libraries: OpenHGNN51, XGBoost, Scikit-learn52, Scikit-optimize53, Captum54, MLstatkit, and Imbalanced-learn55 were used.
The MedHG-PS was compared with traditional ML models, including three representative ML algorithms: logistic regression (LR), multi-layer perceptron (MLP), and XGBoost56. These methods were evaluated using demographic and perioperative information alone, as well as in combination with different network science features to analyze provider interactions and patient transfers within the hospital.
To obtain network science features, the provider information of all hospital encounters was constructed as a single holistic provider-to-provider interaction graph, where nodes are individual providers and undirected edge, and the weights represent the number of interactions—two providers operated on the same patient. Then, the inpatient transfer information of each single hospital encounter was modeled as individual, standalone patient transfer graphs, where nodes represented hospital care units and directed edges indicated patient transfers between units (e.g., a patient from an acute care unit to an intensive care unit) during a single hospital stay. Various network metrics57,58 such as centralities, average degree connectivity, clustering coefficient, authority, page rank, density, number of nodes, and weighted number of edges were extracted from these graphs. For each encounter, these metrics were computed for each node (providers in the provider-to-provider interaction graph and care units in the patient transfer graphs) and then aggregated them into summary statistics, including maximum, minimum, mean, median, and interquartile range. As a result, the inputs for ML models included encounter-level demographics, perioperative information, and graph metrics derived from provider-to-provider interaction and patient transfer graphs.
Following ML best practices, the dataset was divided into training, validation, and testing sets with a ratio of 8:1:1 for model training and hyperparameter tuning. The detailed search space for hyperparameter tuning for different methods is summarized in Table 2. For the ie-HGCN in MedHG-PS, the Adam optimizer59 with binary cross-entropy loss (Eq. (4)) was used to train the MedHG-PS and tree-structured parzen estimator algorithm60 was used to optimize the learning rate, L2 regularization, dropout rate, batch normalization, the dimension of the first hidden layer, and attention vector. The training batch size was set to 51261.
For the ML baselines, Bayes algorithm62 was used to tune hyperparameters and optimize performance. An early stopping strategy was applied to all training processes by monitoring the AUROC changes on the validation set. For the XGBoost, the L2 regularization was set to 1, and the L1 regularization was set to 0. For the MLP, the number of layers was set to 3, and the activation layers were ReLU63. The MLP was trained for a maximum of 200 iterations with the Adam optimizer with cross-entropy loss in an initial learning rate of 0.001. The training data was divided into mini-batches for MLP training, and the mini-batch size was 200. The LR models were optimized using the Newton-CG algorithm with L2 regularization with 500 iterations.
To evaluate the effectiveness of the imputation strategy, a masking approach was implemented based on previous studies. Prior research64,65,66 suggests that demographic information is generally well-recorded in EHR, with the exception of BMI, which may be missing at a rate of approximately 1.7% to 4.3%67,68, primarily due to the absence of height data. Marriage status data may also be missing at a rate of around 2.11%64. In addition, CCI might have a higher missing rate to 8.3%69. In this experiment, BMI and height were randomly masked at a rate of 4.3%, while marital status was masked at a rate of 2.11%. CCI was also masked at a rate of 8.3%. Given the advancements in EHR systems, most procedural intervals and calendar-related factors are recorded seamlessly. In addition, it remains challenging to distinguish between missing and truly negative for comorbidities, the presence of delirium, and the occurrence of anesthesia. Therefore, these features were not masked. Additionally, ASA-PS was masked at a rate of 1.7, consistent with previous findings68. The masked features were then imputed using the proposed methods.
All the models were evaluated on the testing set. Six typical metrics were adopted to assess model performance, including AUROC70, AUPRC71, F1-score, precision, recall72, and specificity73. To examinate the statistical significance between AUROC of different models, DeLong’s test74 was applied. 95% CI for AUROC and AUPRC were estimated using bootstrapping75.
link





More Stories
OhioHealth Physician Group Primary Care Earns American Heart Association Awards
Aspirus Stevens Point Hospital Receives 2026 WHA Quality and Patient Safety Award | Press Room
SGMC Health Wins Two Statewide Patient Safety Awards from the Georgia Hospital Association